P9: Proteases for essential lysosome functions
Cathepsin proteases in inherited disease and infection (PI Thomas Reinheckel)
Cathepsins are lysosomal proteases generally considered to be redundant enzymes that mediate destructive protein breakdown in the acidic milieu of endosomes and lysosomes. Recessive loss-of-function mutations in six of the fifteen cathepsin enzymes encoded in the human genome are causative for clinically very distinct inherited diseases. Some of this diversity might be explained by differential expression of cathepsins in certain cell types, but much of it is likely to be determined by the activity of substrate delivery to the lysosome, i.e. by phagocytosis, endocytosis or autophagy, as well as by intrinsic biochemical cleavage selectivity of the individual cathepsins. A key question will be whether there are indeed specific cleavages consistently attributable to a specific cathepsin as some literature suggests.
ProtPath offers two topics for doctoral work on cathepsins:
1) Cathepsin D substrates and Neurodegeneration
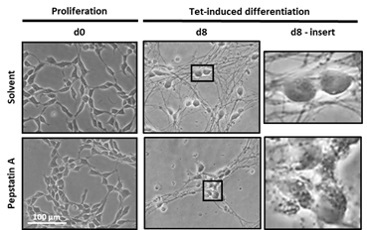
Using the LUHMES cell model we will address neuronal ceroid lipofuscinosis type 10 (CLN10), which is caused by defective cathepsin D. We use proteome analyses to determine cathepsin D substrates and validate those candidates by genetic, biochemical and cell biological methods.
2) Cathepsin substrates in bacteria-infected macrophages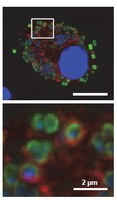
in the phagolysosome (red). Staining for nucleic acids (blue).
Cathepsins reside in the phagolysosome and contribute to the killing of bacteria – but it is unknown how they accomplish this.
Using macrophages as cell model we will determine the S. aureus and macrophage proteins that are cleaved by cathepsin proteases. We further validate and functionally analyse those candidates by genetic, biochemical and cell biological methods.