P5: Deubiquinating enzymes in cell regulation
Regulatory roles of deubiquitinating enzymes (DUBs) (PI Klaus-Peter Knobeloch)
Deubiquitinating enzymes (DUBs) counteract ubiquitin- modification and thus regulate pleiotropic cellular functions including protein stability, function and complex formation, endocytosis, cell signaling, autophagy, subcellular localisation and DNA repair. Research in the field is not only important to define the molecular role of particular DUBs but is also a prerequisite to estimate consequences of DUB inhibition by small molecules. These therapeutic approaches gain growing interest boosted by the development of selective inhibitors showing promising results for the treatment of various human diseases.
ProtPath offers two topics for doctoral work on deubiquitinating enzymes:
Analyzing the molecular and physiological function of USP15 in vivo
USP15 is a DUB which was shown to play a fundamental role in neuroinflammation and mitophagy. To further define physiological and molecular functions, we have generated a conditional k.o mouse model for USP15. Using this model system, the role of USP15 in physiological processes will be defined. Furthermore, in vivo substrates will be identified by comparing the ubiquitin-modified proteome between wildtype and USP15 deficient cells/ organs using a proteomics approach depicted in figure 1. The overall aim is to gain insight into the molecular and functional role of this ubiquitin–specific protease in health and disease.
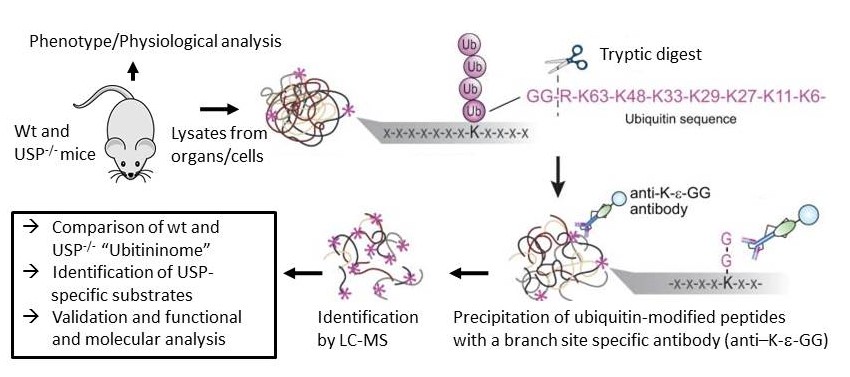
Ubiquitinome will be profiled by the Ubiscan method which is established in our lab. Antibodies detecting the diGly motif characteristic for Ub modified proteins are used to enrich respective peptides after tryptic digest. Subsequently these sites are identified by LC-MS analysis.
Redundant and specific functions of the highly related DUBs USP4, USP15 and USP11
USP15 exhibits considerable sequence similarity and highly related domain structure with USP4 and USP11 (Fig.2). This highly related subfamily of DUBs thus constitutes an excellent research subject to decipher principles of substrate recognition, physiological relevance, cell specificity, redundancy and structural-functional relationships.The project will employ mouse genetics and CRISPR/Cas9 technology to dissect overlapping and redundant physiological and molecular functions. The overall aim will be to define structural-functional relationships of substrate recognition and evaluate consequences and therapeutic options of small molecule inhibition targeting members of the USP15/4/11 subfamily.
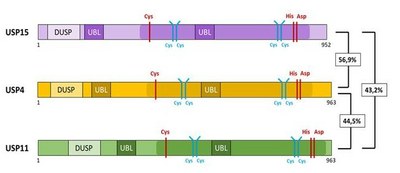
Fig. 2: Domain Structure of USP15, USP4 and USP11.
USP15, 11 and USP4 represent highly related Ubiquitin isopeptidases with a similar domain structure, size and remarkable homology as indicated by percentages of aminoacid (AA) identity. (Vlasschaert et al. 2015). AA forming the catalytic triad are marked in red. UBL: ubiquitin like domain; DUSP: domain present in ubiquitin-specific proteases.

